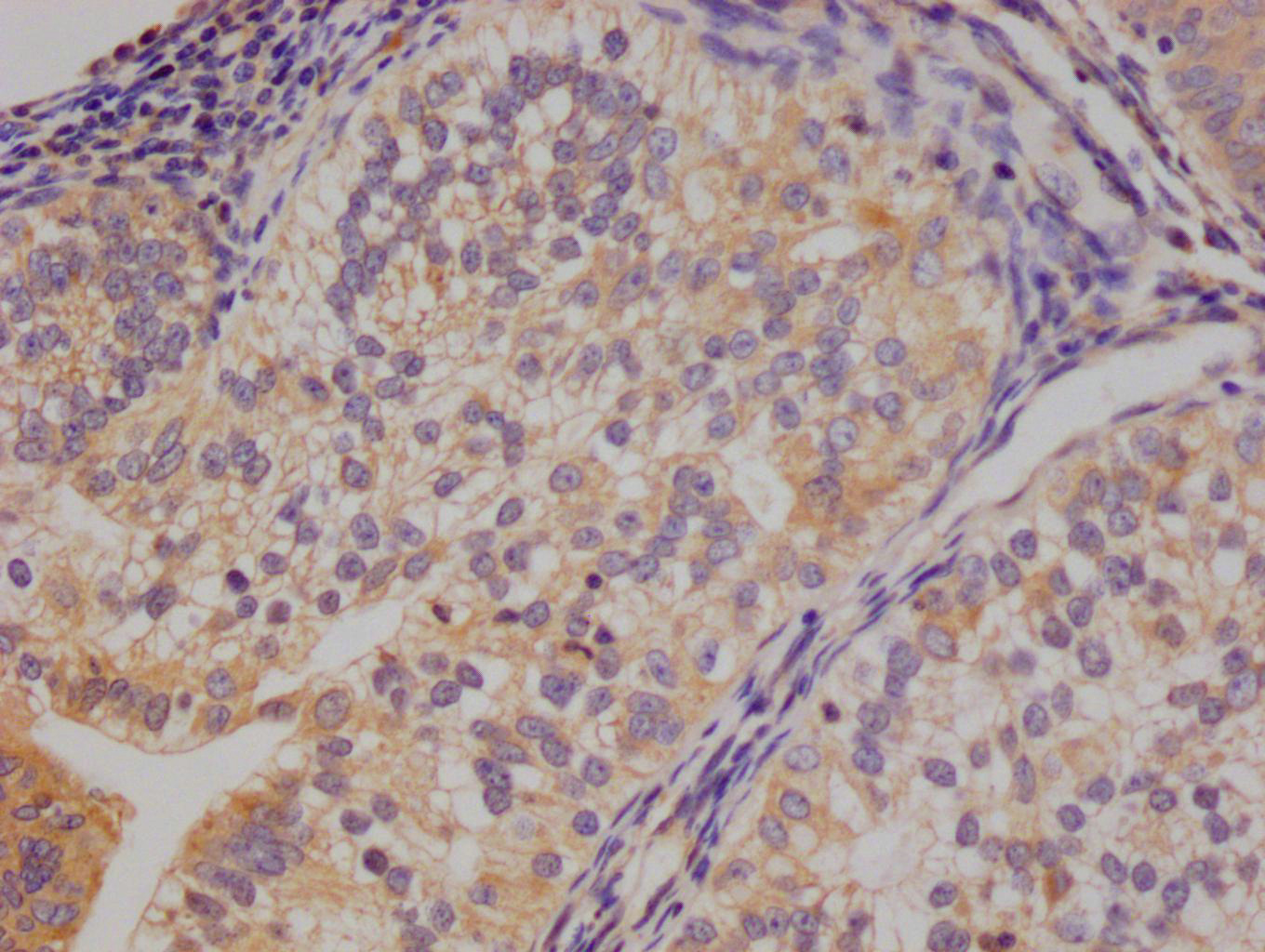
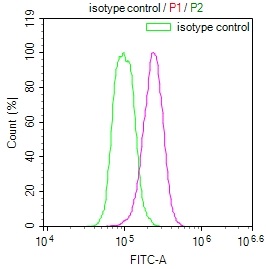
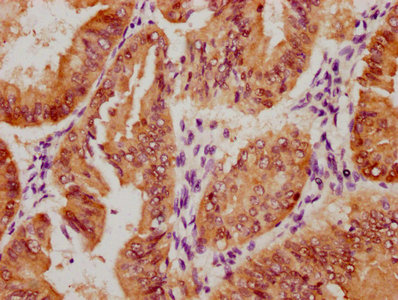
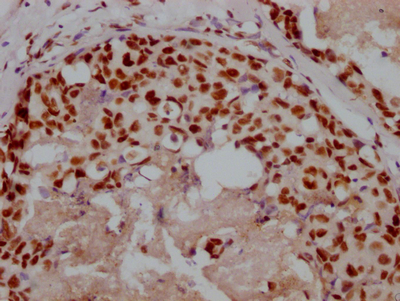
ADAR Antibody, FITC conjugated
-
中文名稱:ADAR兔多克隆抗體, FITC偶聯
-
貨號:CSB-PA001324LC01HU
-
規格:¥880
-
其他:
產品詳情
-
產品名稱:Rabbit anti-Homo sapiens (Human) ADAR Polyclonal antibody
-
Uniprot No.:
-
基因名:
-
別名:136 kDa double-stranded RNA-binding protein antibody; 136kDa double stranded RNA binding protein antibody; Adar 1 antibody; ADAR antibody; Adar1 antibody; Adenosine deaminase acting on RNA 1 A antibody; Adenosine deaminase RNA specific 1 antibody; Adenosine deaminase RNA specific antibody; Adenosine deaminase that act on RNA antibody; AGS6 antibody; AV242451 antibody; Double stranded RNA specific adenosine deaminase antibody; Double-stranded RNA-specific adenosine deaminase antibody; Double-stranded RNA-specific editase Adar antibody; DRADA antibody; Dsh antibody; Dsrad antibody; DSRAD_HUMAN antibody; dsRNA adenosine deaminase antibody; EC 3.5.4.- antibody; G1P1 antibody; IFI 4 antibody; IFI-4 antibody; IFI4 antibody; Ifi4 protein antibody; Interferon induced protein 4 antibody; Interferon inducible protein 4 antibody; Interferon-inducible protein 4 antibody; K88DSRBP antibody; mZaADAR antibody; P136 antibody; Pre-mRNA adenosine deaminase antibody; RNA adenosine deaminase 1 antibody; RNA-editing deaminase 1 antibody; RNA-editing enzyme 1 antibody
-
宿主:Rabbit
-
反應種屬:Human
-
免疫原:Recombinant Human Double-stranded RNA-specific adenosine deaminase protein (367-471AA)
-
免疫原種屬:Homo sapiens (Human)
-
標記方式:FITC
-
克隆類型:Polyclonal
-
抗體亞型:IgG
-
純化方式:>95%, Protein G purified
-
濃度:It differs from different batches. Please contact us to confirm it.
-
保存緩沖液:Preservative: 0.03% Proclin 300
Constituents: 50% Glycerol, 0.01M PBS, pH 7.4 -
產品提供形式:Liquid
-
儲存條件:Upon receipt, store at -20°C or -80°C. Avoid repeated freeze.
-
貨期:Basically, we can dispatch the products out in 1-3 working days after receiving your orders. Delivery time maybe differs from different purchasing way or location, please kindly consult your local distributors for specific delivery time.
-
用途:For Research Use Only. Not for use in diagnostic or therapeutic procedures.
相關產品
靶點詳情
-
功能:Catalyzes the hydrolytic deamination of adenosine to inosine in double-stranded RNA (dsRNA) referred to as A-to-I RNA editing. This may affect gene expression and function in a number of ways that include mRNA translation by changing codons and hence the amino acid sequence of proteins; pre-mRNA splicing by altering splice site recognition sequences; RNA stability by changing sequences involved in nuclease recognition; genetic stability in the case of RNA virus genomes by changing sequences during viral RNA replication; and RNA structure-dependent activities such as microRNA production or targeting or protein-RNA interactions. Can edit both viral and cellular RNAs and can edit RNAs at multiple sites (hyper-editing) or at specific sites (site-specific editing). Its cellular RNA substrates include: bladder cancer-associated protein (BLCAP), neurotransmitter receptors for glutamate (GRIA2) and serotonin (HTR2C) and GABA receptor (GABRA3). Site-specific RNA editing of transcripts encoding these proteins results in amino acid substitutions which consequently alters their functional activities. Exhibits low-level editing at the GRIA2 Q/R site, but edits efficiently at the R/G site and HOTSPOT1. Its viral RNA substrates include: hepatitis C virus (HCV), vesicular stomatitis virus (VSV), measles virus (MV), hepatitis delta virus (HDV), and human immunodeficiency virus type 1 (HIV-1). Exhibits either a proviral (HDV, MV, VSV and HIV-1) or an antiviral effect (HCV) and this can be editing-dependent (HDV and HCV), editing-independent (VSV and MV) or both (HIV-1). Impairs HCV replication via RNA editing at multiple sites. Enhances the replication of MV, VSV and HIV-1 through an editing-independent mechanism via suppression of EIF2AK2/PKR activation and function. Stimulates both the release and infectivity of HIV-1 viral particles by an editing-dependent mechanism where it associates with viral RNAs and edits adenosines in the 5'UTR and the Rev and Tat coding sequence. Can enhance viral replication of HDV via A-to-I editing at a site designated as amber/W, thereby changing an UAG amber stop codon to an UIG tryptophan (W) codon that permits synthesis of the large delta antigen (L-HDAg) which has a key role in the assembly of viral particles. However, high levels of ADAR1 inhibit HDV replication.
-
基因功能參考文獻:
- This study investigates the genome-wide binding preferences of the nuclear constitutive isoforms ADAR1-p110 and ADAR2 on human miRNA species by RNA immunoprecipitation of ADAR-bound small RNAs . PMID: 29950133
- The adenosine deaminase RNA-specific binding protein (ADAR1)-dependent and RNA-editing-independent regulation of invasion, mediated by Integrin beta3 (ITGB3) suggests a central involvement of ADAR1 in cancer progression and metastasis. PMID: 29855470
- The c.3463C>T missense mutation of the ADAR1 gene probably underlies the dyschromatosis symmetrical hereditaria in the Chinese pedigree. PMID: 29896739
- These findings suggest that Zalpha domain of human ADAR1 binding with the GAC hairpin stem in COMP can lead to a non-genetic, RNA editing-mediated substitution in COMP that may then play a crucial role in the development of pseudoachondroplasia. PMID: 28924040
- Expression of wild-type, but not mutant, ADAR1 enhances Alu-dependent editing and transcriptional activity of GLI1, a Hedgehog (Hh) pathway transcriptional activator and self-renewal agonist, and promotes immunomodulatory drug resistance in vitro. PMID: 29203771
- ADAR1 silencing resulted in a lower global double-stranded to single-stranded RNA ratio, suggesting that A-to-I editing can stabilize a large subset of imperfect RNA duplexes. The results imply that the editing effect on RNA secondary structure is context dependent and underline the intricate regulatory role of ADAR1 on global RNA secondary structure. PMID: 29129909
- The Adenosine deaminase acting on RNA1 catalyzes the C6 deamination of adenosine (A) to produce inosine (I) in regions of RNA with double-stranded (ds) character. PMID: 24905200
- ADAR copy number is negatively associated with apoptosis and several immune cell types' signatures. PMID: 29273356
- CDK13 RNA over-editing sites mediated by ADAR1 may serve as novel cancer driver events in HCC progression. PMID: 29996118
- n this study, we found that a 28 year-old male patient harbouring a deleterious substitution of Leu1052Pro in the ADAR1 gene in a typical Dyschromatosis symmetrica hereditaria family. His mother suffered from the Dyschromatosis symmetrica hereditaria also owns the same mutation. PMID: 29321362
- ADAR1 regulates post-transcriptional gene expression in gastric cancer through both RNA editing-dependent and editing-independent mechanisms. PMID: 29691780
- Expansion of mutation spectrum in dyschromatosis symmetrica hereditaria by identifying eight novel ADAR1 mutations. PMID: 29185800
- ADAR1 functions other than RNA editing are reviewed. PMID: 28960389
- A novel function of ADAR1 as an inhibitor of L1 retrotransposition was demonstrated. PMID: 28640667
- Data collected from our center strongly suggest that ADAR1 expression can effectively predict HCC patients' prognosis and an abnormal overexpression of ADAR1 is positively correlated with AR in HCC. PMID: 29144509
- Results implicate rare variants in the Aicardi-Goutieres syndrome genes ADAR and RNASEH2B and a type I interferon signature in glioma and prostate carcinoma risk and tumorigenesis, consistent with a genetic basis underlying inflammation-driven malignant transformation in glioma and prostate carcinoma development. PMID: 29030706
- high-throughput mutagenesis analysis (Sat-FACS-Seq) of conserved residues in an RNA-binding loop of hADAR1d revealed essential amino acids for function, advancing our understanding of RNA recognition by this domain PMID: 29457714
- The RNA-editing enzyme ADAR promotes lung adenocarcinoma migration and invasion by stabilizing FAK. PMID: 28928239
- Study investigated genetic, phenotypic, and interferon status of 46 patients from 37 families with neurological disease due to mutations in ADAR1. Phenotype encompassed a spectrum of Aicardi-Goutieres syndrome, isolated bilateral striatal necrosis, spastic paraparesis with normal neuroimaging, a progressive spastic dystonic motor disorder, and adult-onset psychological difficulties with intracranial calcification. PMID: 28561207
- the present study performed a mutation analysis of the ADAR1 gene in two sporadic patients with typical DSH from birth, and identified two novel mutations. PMID: 28393185
- ADAR1 contributes to gastric cancer development and progression via activating mTOR/p70S6K/S6 ribosomal protein signaling axis. PMID: 27863387
- These results extended the known functions of ADAR1 and RNA editing to the critical fine-tuning of the intracellular apoptotic signaling and also provided mechanistic explanation for ADAR1's roles in development and tumorigenesis. PMID: 28542129
- These results demonstrate that ablation of RNase L activity promotes survival of ADAR1 deficient cells even in the presence of MDA5 and MAVS, suggesting that the RNase L system is the primary sensor pathway for endogenous dsRNA that leads to cell death. PMID: 28362255
- discuss the function of ADAR1 and its regulatory role in viral infection, and establish the relationship between ADAR1 and virus-associated sepsis (review) PMID: 28199207
- The range of human disease associated with ADAR1 mutations may extend further to include other inflammatory conditions while ADAR2 mutations may affect psychiatric conditions. PMID: 28913566
- ADAR1 is targeted by miR-143 to regulate IL-1beta-induced HUVEC activation PMID: 28527816
- The miRNAs that were found upregulated in DENV-infected cells did not control the production of infectious virus particles. On the other hand, miR-3614-5p, which was upregulated in DENV-negative macrophages, reduced DENV infectivity and regulated ADAR1 expression, a protein that facilitates viral replication. PMID: 29045406
- We detected high ADAR1 expression in half of the triple-negative breast cancer patients. In addition to DNA mutations, RNA editing can be related to neoantigens; hence, we need to explore non-synonymous mutations exclusively found using RNA sequencing data to identify clinically relevant neoantigens. PMID: 29022489
- Editing in the STAT3 intron is performed by ADAR1 and affects STAT3 alternative splicing. RNA editing is one of the molecular mechanisms regulating the expression of STAT3beta. PMID: 28278381
- a role for ADAR1 as a lung cancer oncogene undergoing gene amplification-associated activation that affects downstream RNA editing patterns and patient prognosis. PMID: 26640150
- In endothelial cells, ADAR1 overexpression induces CTSS RNA editing and consequently increases cathepsin S expression. ADAR1 levels and the extent of CTSS RNA editing are associated with changes in cathepsin S levels in patients with atherosclerotic vascular diseases. PMID: 27595325
- the activation of the JAK/STAT pathway is a regulatory mechanism of ADAR1 expression and causes abnormal RNA editing profile in esophageal squamous cell carcinoma. This mechanism may serve as a new target for esophageal squamous cell carcinoma therapy PMID: 28714361
- Lentiviral ADAR1 wild-type, but not an editing-defective ADAR1(E912A) mutant, induces self-renewal gene expression and impairs biogenesis of stem cell regulatory let-7 microRNAs. PMID: 27292188
- ADAR1p110 isoform competitively inhibits binding of Staufen1 to the 3'-untranslated-region dsRNAs and antagonizes Staufen1-mediated mRNA decay. PMID: 28436945
- we have identified two novel mutations of the ADAR1 gene and explored their implications on the structure and functions of ADAR1 protein. PMID: 27230815
- ADAR1 restricts LINE-1 retrotransposition. PMID: 27658966
- It was established in this report that interactions between PACT, ADAR1 and HIV-1-encoded Tat protein diminish the activation of PKR in response to HIV-1 infection. PMID: 28167698
- Studies suggest that the overexpression of ADAR1 protein forms a feedback loop starting with a decreased microRNA miR-17-5p level, that in turn up-regulates ADAR1. PMID: 28398248
- both lysine 574 and 576 are essential for ADAR1-P110 ubiquitination. PMID: 27729454
- The molecular effects of ADAR1 overexpression in HEK293T cells by label-free quantitative proteomics has been reported. PMID: 27104882
- A novel mutation in exome 8 of ADAR1 was identified (c.2633-2634delCT) in all the affected members of family 1 (Fig. 1h). A novel mutation in exome 2 of ADAR1 was also identified (c.1057-1058delTG) in all the affected members of family 2 (Fig. 2f). These two mutations were not reported before and not found in the unaffected individuals of the families and 100 unrelated Chinese controls PMID: 25763870
- This study reviewed that Neurologic Phenotypes Associated with Mutations in ADAR1 in patients with Aicardi-Goutieres Syndrome. PMID: 27643693
- Results indicated that ADAR1 might play an important role in the occurrence, progression and prognosis of cervical squamous cancer PMID: 28109322
- These findings suggest that adenosine deaminase acting on RNA 1 is subject to different regulations by DNA methyltransferase and histone deacetylase enzymes in neuronal SH-SY5Y cells. PMID: 26485095
- ADAR1 p150 as the major A-to-I editor in mouse embryo fibroblasts. PMID: 26817845
- ADAR1, in an editing-independent manner, regulates the biogenesis of miR-222 at the transcription level and thereby ICAM1 expression, which consequently affects melanoma immune resistance. PMID: 26338962
- The frame-shift mutation c.2638delG and the nonsense mutation c.2867C>A in the ADAR1 gene are associated with the dyschromatosis symmetrica hereditaria. PMID: 27060309
- Recent research has demonstrated that inosine in the epitranscriptome and ADAR1 protein establish innate immune tolerance for host dsRNA formed by endogenous sequences. PMID: 26658668
- A-to-I RNA editing levels catalyzed by ADAR1 and ADARB1 decreased in Alzheimer's disease patients' brain tissues, mainly in the hippocampus and to a lesser degree in the temporal and frontal lobes. PMID: 26655226
- A three-generation family exhibiting phenotypic variability with a single germline ADAR1 mutation suggests that chilblain might aggravate the clinical phenotypes of dyschromatosis symmetrica hereditaria. PMID: 26892242
顯示更多
收起更多
-
相關疾病:Dyschromatosis symmetrica hereditaria (DSH); Aicardi-Goutieres syndrome 6 (AGS6)
-
亞細胞定位:[Isoform 1]: Cytoplasm. Nucleus.; [Isoform 5]: Cytoplasm. Nucleus. Nucleus, nucleolus.
-
組織特異性:Ubiquitously expressed, highest levels were found in brain and lung. Isoform 5 is expressed at higher levels in astrocytomas as compared to normal brain tissue and expression increases strikingly with the severity of the tumor, being higher in the most ag
-
數據庫鏈接:
Most popular with customers
-
-
YWHAB Recombinant Monoclonal Antibody
Applications: ELISA, WB, IHC, IF, FC
Species Reactivity: Human, Mouse, Rat
-
Phospho-YAP1 (S127) Recombinant Monoclonal Antibody
Applications: ELISA, WB, IHC
Species Reactivity: Human
-
-
-
-
-